Lower Duncan River Kokanee AUC Analysis 2016
The suggested citation for this analytic report is:
Thorley, J.L. (2016) Lower Duncan River Kokanee AUC Analysis 2016. A Poisson Consulting Analysis Report. URL: https://www.poissonconsulting.ca/f/1172999120.
Background
Every fall, Kokanee from Kootenay Lake spawn in the Duncan-Lardeau system. Since 2008 aerial surveys have been conducted in the Lower Duncan River (LDR) to enumerate the number of kokanee spawners. The management questions are:
- What is the spawn run timing, fry emergence timing, and relative intensity of kokanee spawning in the Lower Duncan River? What potential operational/environmental cues affect this variable?
- What are the timing/cues of kokanee spawners in Meadow Creek and Lardeau River systems?
- What are the relative distribution of kokanee spawners in the Lower Duncan River, Meadow Creek and Lardeau River? What potential operation/environmental/physical cues (e.g., temperature, velocity, depth, cover, substrate) affect this variable?; and
- What physical works or operational constraints could be implemented to minimize operational conflicts associated with recommended kokanee spawning operations?
Methods
Data Preparation
The survey data were provided by LGL Limited in the form of an Excel database. The referential integrity of the data table’s relationships was confirmed using SQL. The mean daily discharge and water temperature values at Duncan River below Lardeau (DRL) were extracted from BC Hydro’s environmental database for the Kootenay’s, which is maintained by Poisson Consulting. The data were clean and tidied (Wickham 2014) using R version 3.3.2 (R Core Team 2016). When there were repeated counts for a channel the higher of the two values was choosen.
Statistical Analysis
Hierarchical Bayesian models were fitted to the count data using R version 3.3.2 (R Core Team 2016) and JAGS 4.2.0 (Plummer 2015) which interfaced with each other via jaggernaut 2.5.1 (Thorley 2013). For additional information on hierarchical Bayesian modelling in the BUGS language, of which JAGS uses a dialect, the reader is referred to (Kery and Schaub 2011, 41–44).
Unless specified, the models assumed vague (low information) prior distributions (Kery and Schaub 2011, 36). The posterior distributions were estimated from a minimum of 1,000 Markov Chain Monte Carlo (MCMC) samples thinned from the second halves of three chains (Kery and Schaub 2011, 38–40). Model convergence was confirmed by ensuring that Rhat (Kery and Schaub 2011, 40) was less than 1.1 for each of the parameters in the model (Kery and Schaub 2011, 61).
The posterior distributions of the fixed (Kery and Schaub 2011, 75) parameters are summarised in terms of a point estimate (median), lower and upper 95% credible limits (2.5th and 97.5th percentiles), the standard deviation (SD), percent relative error (half the 95% credible interval as a percent of the point estimate) and significance (Kery and Schaub 2011, 37, 42).
Variable selection was achieved by dropping insignificant (Kery and Schaub 2011, 37, 42) fixed (Kery and Schaub 2011, 77–82) variables and uninformative random variables. A fixed variables was considered to be insignificant if its significance was \(\geq\) 0.05 while a random variable was considered to be uninformative if its percent relative error was \(\geq\) 80%. The Deviance Information Criterion (DIC) was not used because it is of questionable validity when applied to hierarchical models (Kery and Schaub 2011, 469).
The results are displayed graphically by plotting the modelled relationships between particular variables and the response with 95% credible intervals (CRIs) with the remaining variables held constant. In general, continuous and discrete fixed variables are held constant at their mean and first level values respectively while random variables are held constant at their typical values (expected values of the underlying hyperdistributions) (Kery and Schaub 2011, 77–82). Where informative the influence of particular variables is expressed in terms of the effect size (i.e., percent change in the response variable) with 95% CRIs (Bradford, Korman, and Higgins 2005).
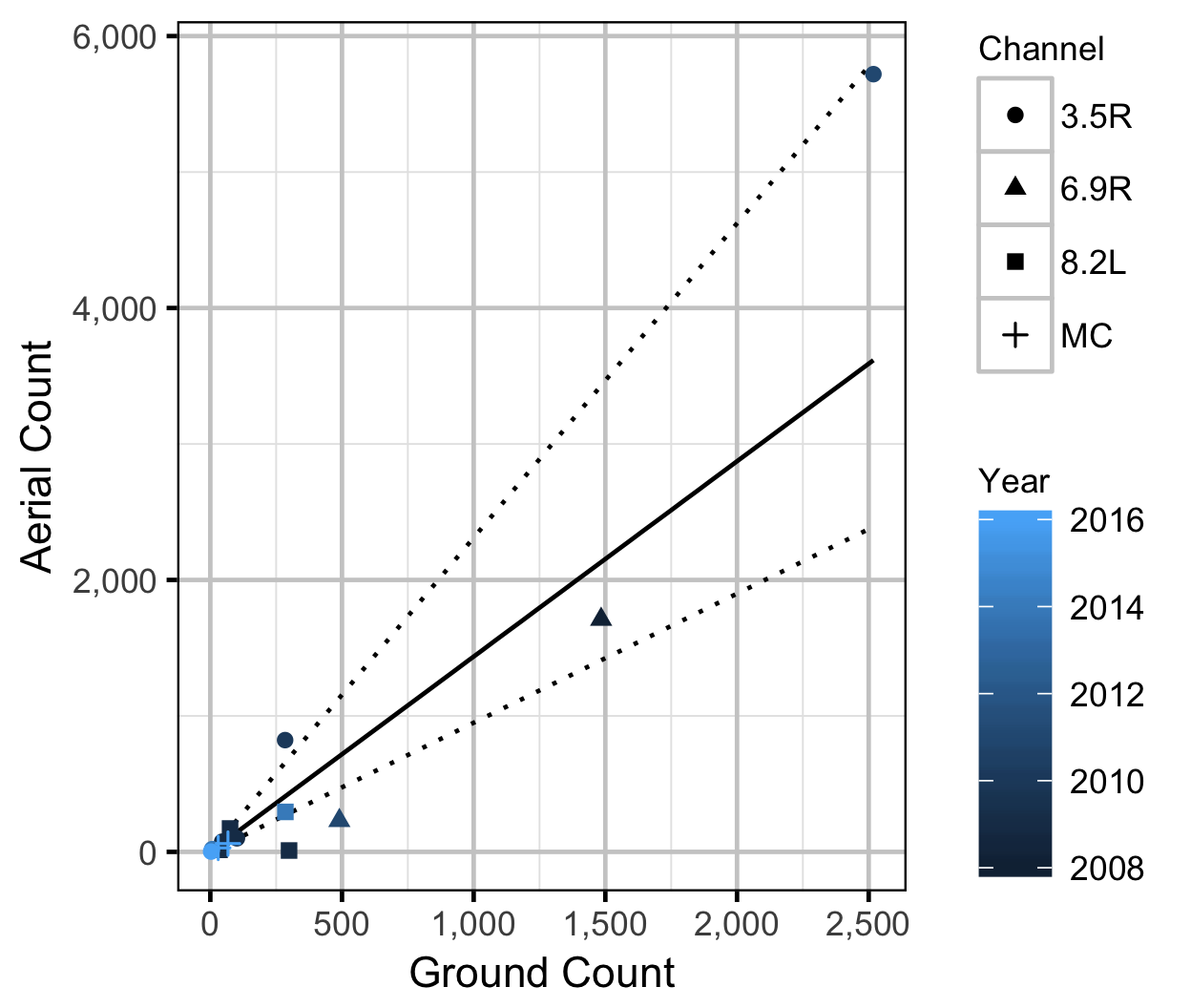
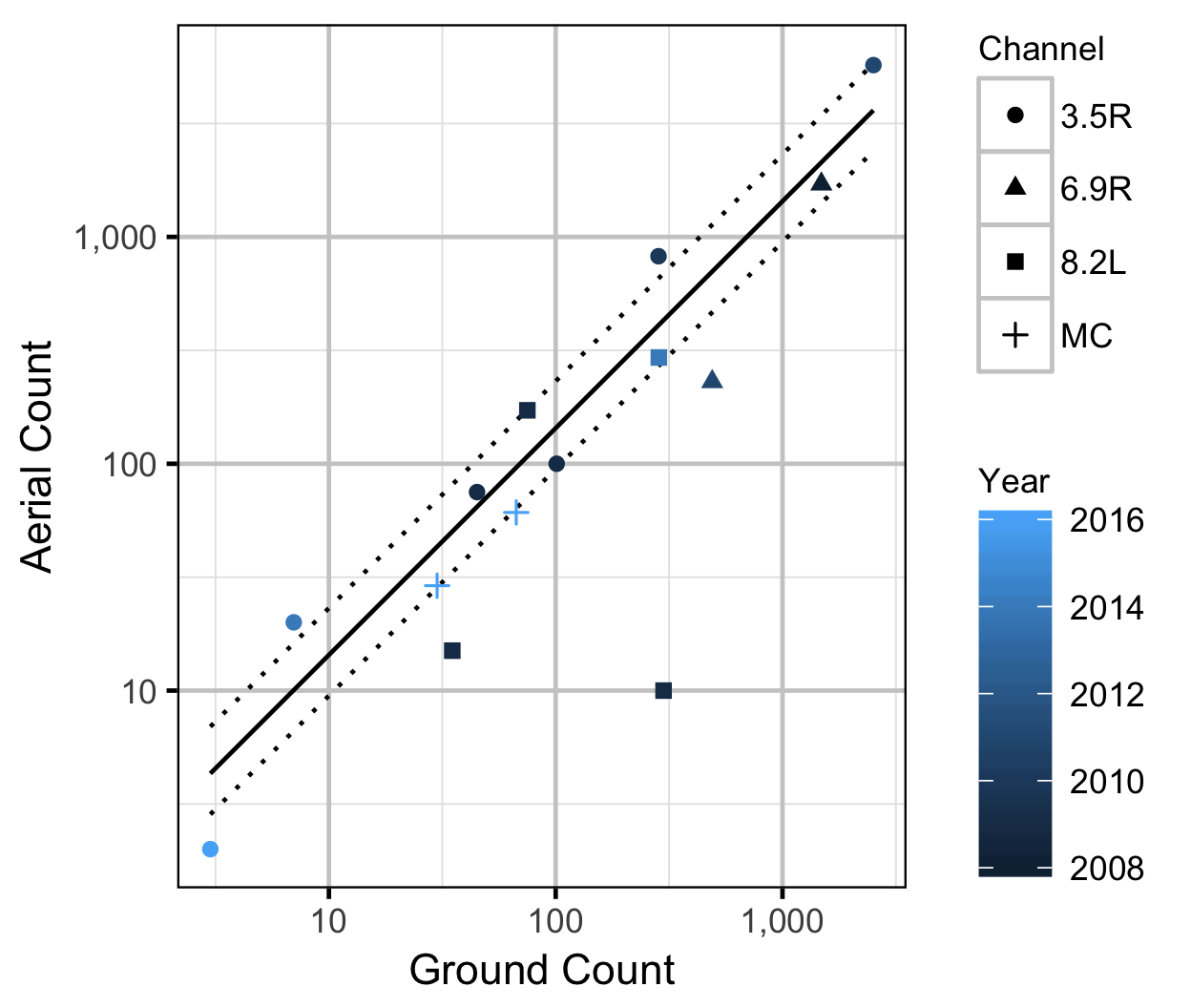
Observer Efficiency
The aerial observer efficiency was estimated from ground counts using an overdispersed Poisson regression. Key assumptions of the observer efficiency model include:
- The ground counts are accurate.
- The residual variation in the aerial counts is gamma-Poisson distributed.
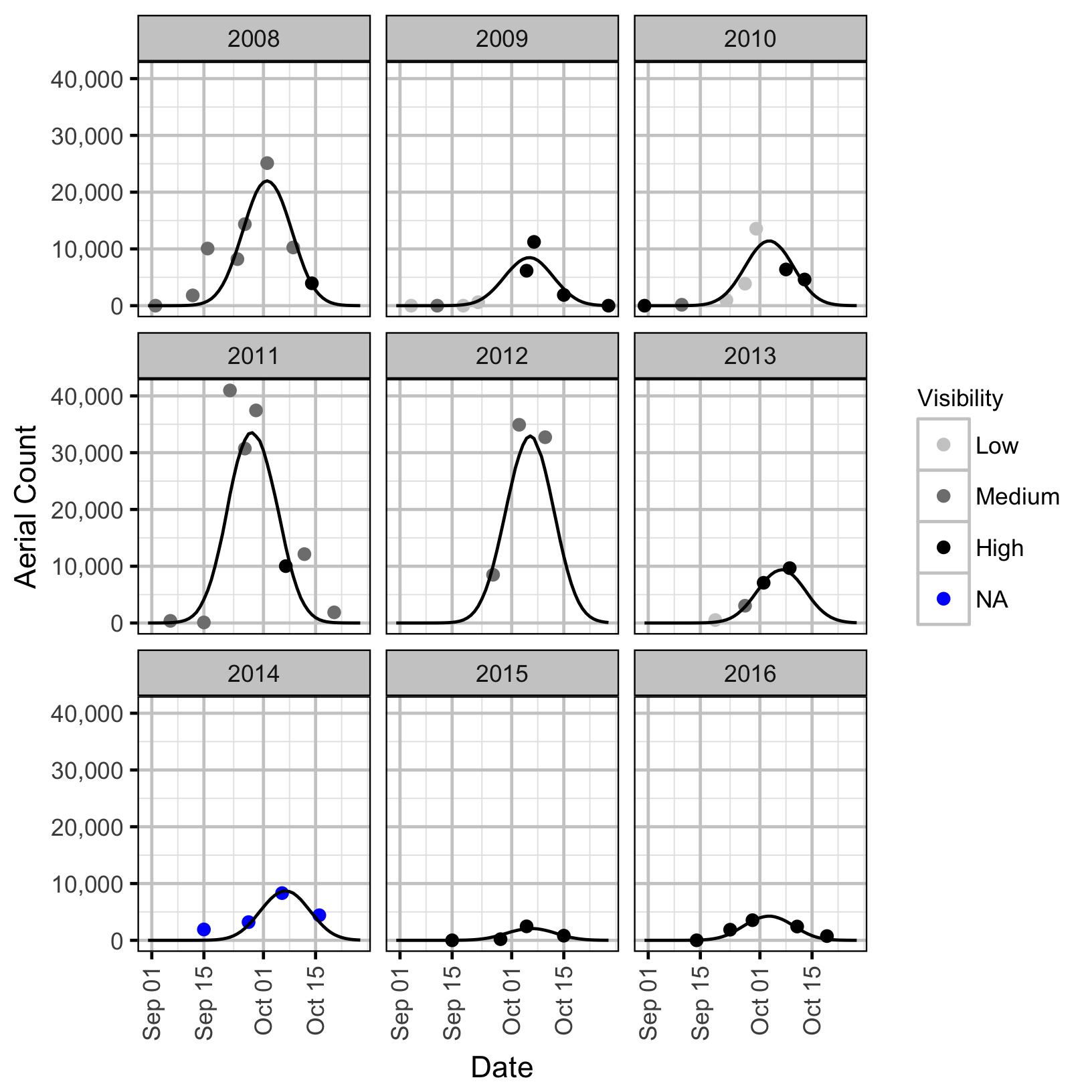
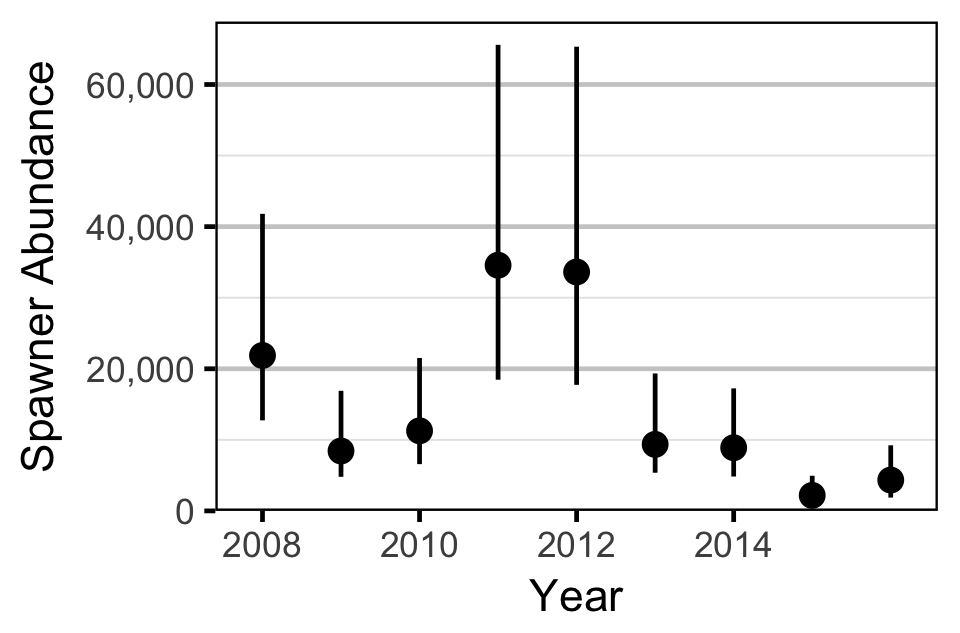
Area-Under-the-Curve
The aerial spawner counts for the Lower Duncan River and its sidechannels were analysed using a hierarchical Bayesian Area-Under-the-Curve (AUC) model (Hilborn, Bue, and Sharr 1999). Key assumptions of the AUC model include:
- Spawner arrival and departure times are normally distributed (Hilborn, Bue, and Sharr 1999).
- Spawner duration is constant across years.
- Spawner abundance varies randomly by year.
- Peak spawn timing varies randomly by year (Su, Adkison, and Van Alen 2001).
- Spawner residence time is between 7 and 14 days (Acara 1970, @morbey_timing_2003).
- Spawner observer efficiency is drawn from a normal distribution as estimated by the observer efficiency model, with a mean of 1.43, SD of 0.35 and lower and upper limits of 0.95 and 2.31 respectively.
- The standard deviation of the residual variation in the spawner counts varies by the annual abundance.
- The residual variation in the spawner counts is normally distributed.
The emergence timing was calculated from the daily water temperature at DRL assuming 1,000 Accumulated Thermal Units (ATUs).
Model Code
The JAGS model code, which uses a series of naming conventions, is presented below.
Observer Efficiency
| Variable/Parameter | Description |
|---|---|
Aerial[i] |
Aerial count for ith survey |
bEfficiency |
eEfficiency intercept |
eAerial[i] |
Expected aerial count for ith survey |
Ground[i] |
Ground count for ith survey |
sDispersion |
SD of overdispersion |
Observer Efficiency - Model1
model{
bEfficiency ~ dunif(0, 3)
sDispersion ~ dunif(0, 5)
for (i in 1:length(Aerial)){
eAerial[i] <- Ground[i] * bEfficiency
eDispersion[i] ~ dgamma(1 / sDispersion^2, 1 / sDispersion^2)
Aerial[i] ~ dpois(eAerial[i] * eDispersion[i])
}
}Area-Under-The-Curve
| Variable/Parameter | Description |
|---|---|
bAbundance |
log(eAbundance) intercept |
bAbundanceYear[i] |
Effect of ith year on log(eAbundance) |
bDuration |
eDuration intercept |
bEfficiency |
eEfficiency intercept |
bPeakTiming |
ePeakTiming intercept |
bPeakTimingYear[i] |
Effect of ith year on ePeakTiming |
bResidenceTime |
Spawner residence time |
Count[i] |
Spawner count for ith survey |
Dayte[i] |
Centred day of the year for ith survey |
eAbundance[i] |
Expected annual spawner abundance for ith survey |
eCount[i] |
Expected spawner count for ith survey |
eDuration[i] |
Expected SD of the duration of spawner arrival timing |
eEfficiency[i] |
Expected aerial observer efficiency for ith survey |
ePeakTiming[i] |
Expected timing of annual peak spawner abundance for ith survey |
eSpawners[i] |
Expected number of spawners for ith survey |
FullYear[i] |
Indicates whether seven or more surveys in ith year |
sAbundanceYear |
SD of effect of year on log(eAbundance) |
sCount |
SD of residual variation in eCount |
sPeakTiming |
SD of effect of year on ePeakTiming |
Area-Under-The-Curve - Model1
model{
bAbundance ~ dnorm(10, 5^-2)
bPeakTiming ~ dunif(-14, 14)
bDuration ~ dnorm(2, 1^-2)
bEfficiency ~ dnorm(1.43, 0.35^-2) T(0.95, 2.31)
bResidenceTime ~ dunif(7, 14)
sAbundanceYear ~ dunif(0, 5)
sPeakTiming ~ dunif(0, 14)
for(i in 1:nYear){
bAbundanceYear[i] ~ dnorm(0, sAbundanceYear^-2)
bPeakTimingYear[i] ~ dnorm(0, sPeakTiming^-2)
}
bSD ~ dnorm(0, 10^-2)
bSDAbundance ~ dnorm(0, 5^-2)
for(i in 1:length(Count)){
log(eAbundance[i]) <- bAbundance + bAbundanceYear[Year[i]]
ePeakTiming[i] <- bPeakTiming + bPeakTimingYear[Year[i]]
log(eDuration[i]) <- bDuration
eArrived[i] <- pnorm(Dayte[i], (ePeakTiming[i] - bResidenceTime/2), eDuration[i]^-2)
eDeparted[i] <- pnorm(Dayte[i], (ePeakTiming[i] + bResidenceTime/2), eDuration[i]^-2)
eSpawners[i] <- (eArrived[i] - eDeparted[i]) * eAbundance[i]
eEfficiency[i] <- bEfficiency
log(eSD[i]) <- bSD + bSDAbundance * eAbundance[i]
Count[i] ~ dnorm(eSpawners[i] * eEfficiency[i], eSD[i]^-2)
}
} Results
Model Parameters
The posterior distributions for the fixed (Kery and Schaub 2011 p. 75) parameters in each model are summarised below.
Observer Efficiency
| Parameter | Estimate | Lower | Upper | SD | Error | Significance |
|---|---|---|---|---|---|---|
| bEfficiency | 1.4360 | 0.9490 | 2.3110 | 0.3500 | 47 | 0.001 |
| sDispersion | 0.8655 | 0.6239 | 1.3156 | 0.1783 | 40 | 0.001 |
| Convergence | Iterations |
|---|---|
| 1.02 | 1e+05 |
Area-Under-The-Curve
| Parameter | Estimate | Lower | Upper | SD | Error | Significance |
|---|---|---|---|---|---|---|
| bAbundance | 9.2710000 | 8.1770000 | 10.1920000 | 0.5050000 | 11 | 0.0010 |
| bDuration | 1.6958000 | 1.3403000 | 1.9796000 | 0.1633000 | 19 | 0.0010 |
| bEfficiency | 1.5340000 | 1.0020000 | 2.1500000 | 0.3030000 | 37 | 0.0010 |
| bPeakTiming | 6.2370000 | 2.7140000 | 9.4400000 | 1.6690000 | 54 | 0.0039 |
| bResidenceTime | 10.9590000 | 7.2210000 | 13.8820000 | 1.9500000 | 30 | 0.0010 |
| bSD | 6.6920000 | 5.8100000 | 7.2830000 | 0.3690000 | 11 | 0.0010 |
| bSDAbundance | 0.0000714 | 0.0000312 | 0.0001784 | 0.0000379 | 100 | 0.0010 |
| sAbundanceYear | 1.0330000 | 0.6160000 | 2.1360000 | 0.4110000 | 74 | 0.0010 |
| sPeakTiming | 3.5620000 | 1.4180000 | 8.1050000 | 1.6490000 | 94 | 0.0010 |
| Convergence | Iterations |
|---|---|
| 1.03 | 20000 |
Figures
Observer Efficiency
Area-Under-The-Curve
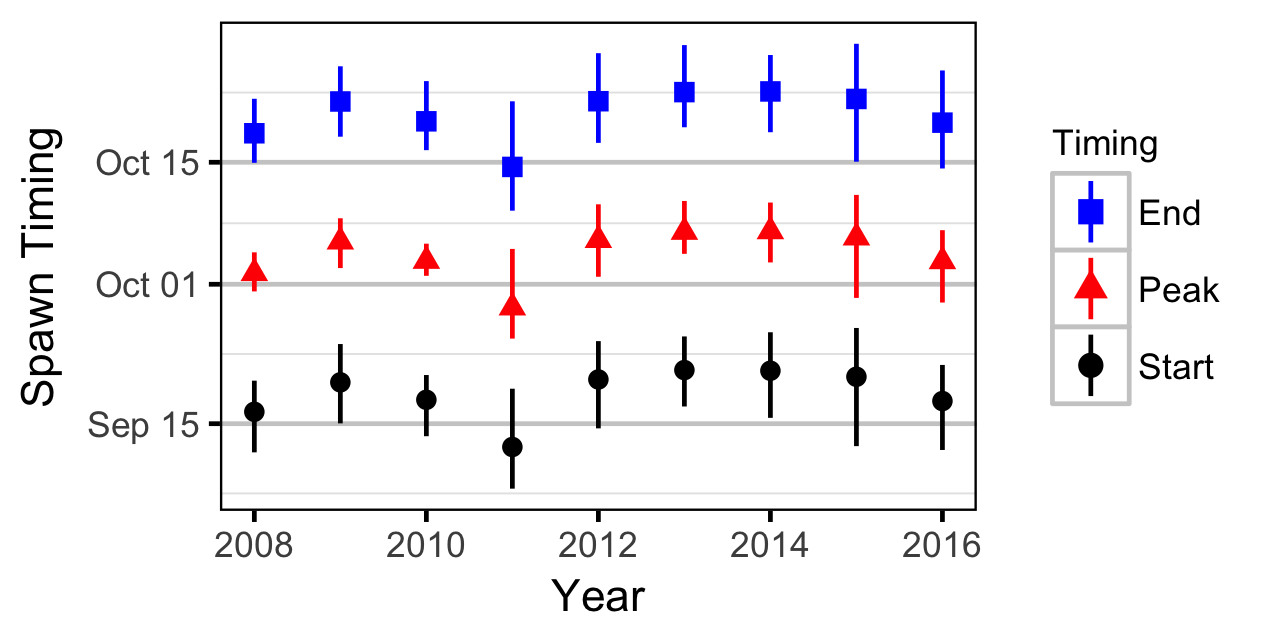
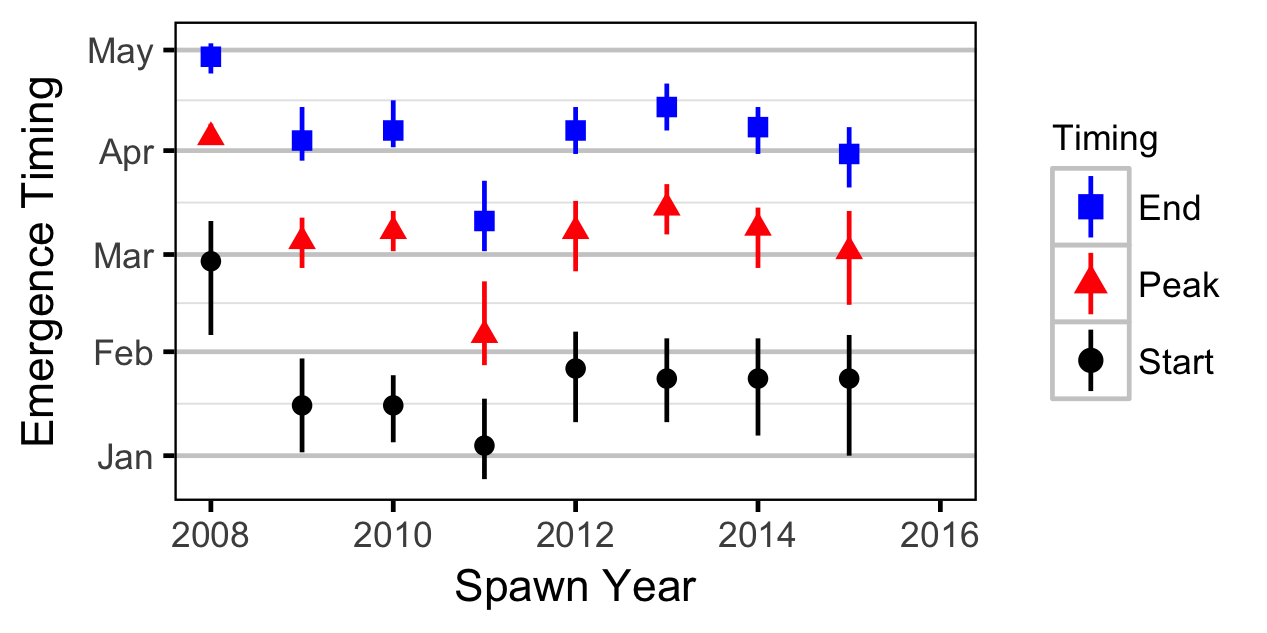
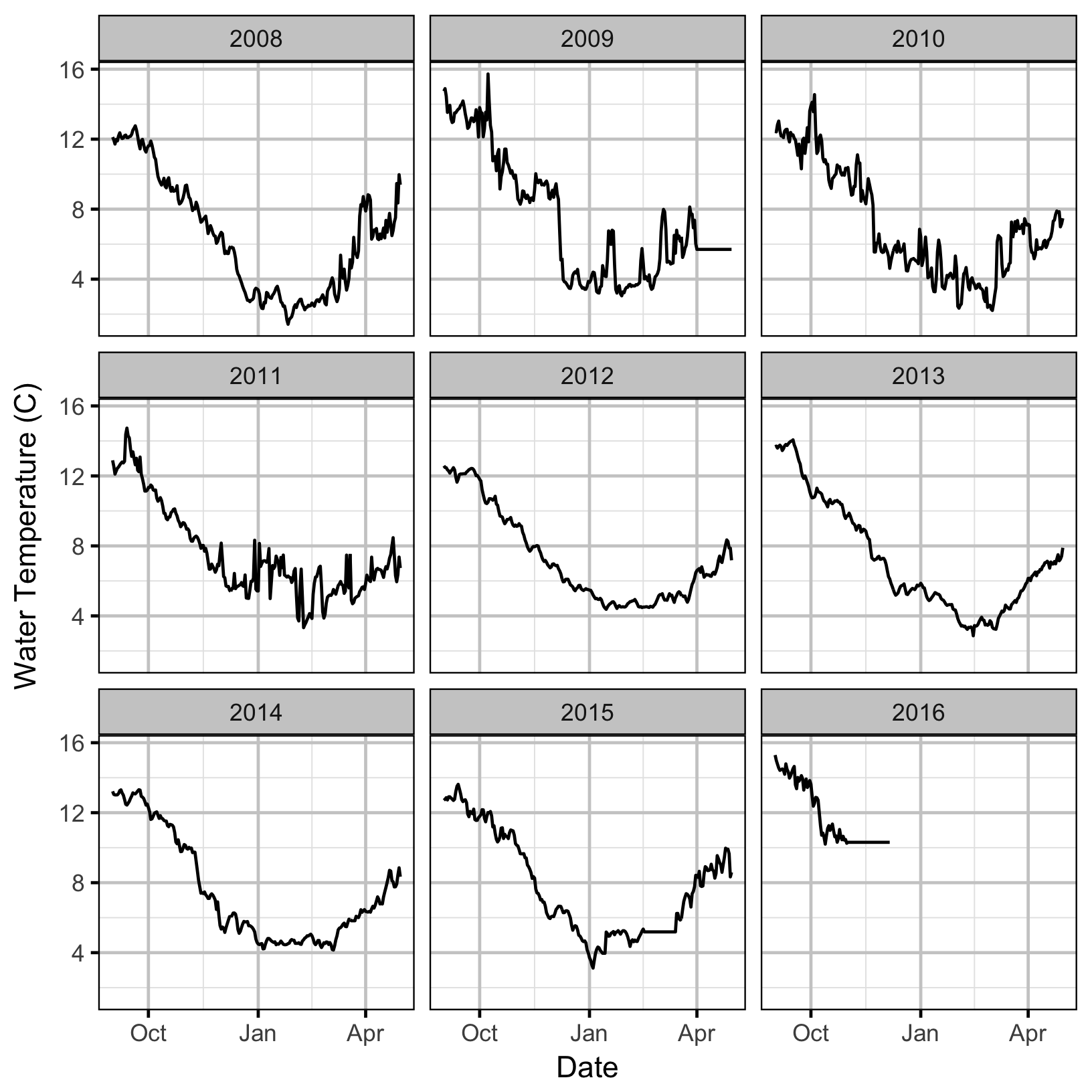
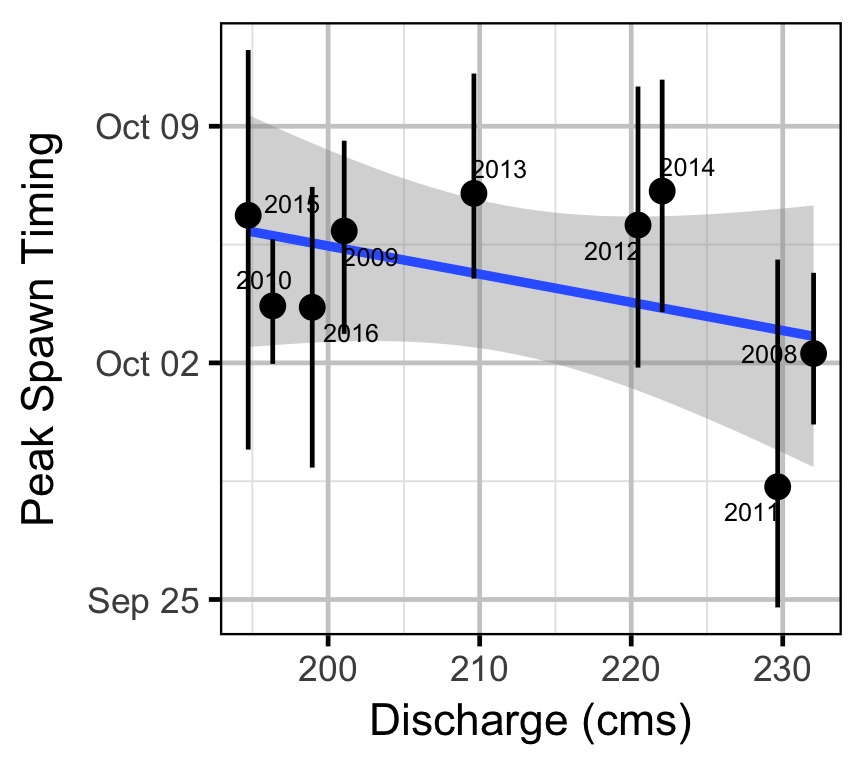
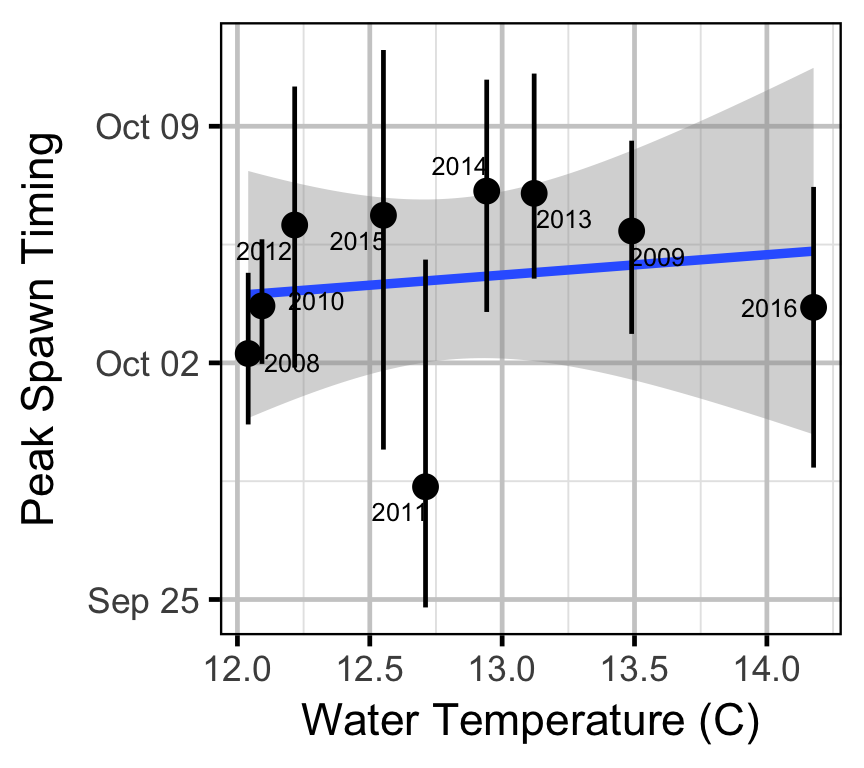
Acknowledgements
The organisations and individuals whose contributions have made this analysis report possible include:
- BC Hydro
- Guy Martel
- Alf Leake
- Margo Sadler
- Phil Bradshaw
- Okanagan National Alliance
- Michael Zimmer
- Amy Duncan
- Natasha Audy
- Skyeler Folks
- Autumn Solomon
- LGL Limited
- Elmar Plate
- Amec Earth and Environmental
- Louise Porto
- Crystal Lawrence
- Poisson Consulting Ltd.
- Robyn Irvine
- Gerry Nellestijn
- Clint Tarala
- Murray Pearson
References
Acara, A. H. 1970. “The Meadow Creek Spawning Channel. Unpublished Report.” Victoria, BC: Fish; Wildlife Branch.
Bradford, Michael J, Josh Korman, and Paul S Higgins. 2005. “Using Confidence Intervals to Estimate the Response of Salmon Populations (Oncorhynchus Spp.) to Experimental Habitat Alterations.” Canadian Journal of Fisheries and Aquatic Sciences 62 (12): 2716–26. https://doi.org/10.1139/f05-179.
Hilborn, Ray, Brian G Bue, and Samuel Sharr. 1999. “Estimating Spawning Escapements from Periodic Counts: A Comparison of Methods.” Canadian Journal of Fisheries and Aquatic Sciences 56 (5): 888–96. https://doi.org/10.1139/f99-013.
Kery, Marc, and Michael Schaub. 2011. Bayesian Population Analysis Using WinBUGS : A Hierarchical Perspective. Boston: Academic Press. http://www.vogelwarte.ch/bpa.html.
Morbey, Y. E., and R. C. Ydenberg. 2003. “Timing Games in the Reproductive Phenology of Female Pacific Salmon (Oncorhynchus Spp.).” The American Naturalist 161 (2): 284–98. http://www.jstor.org/stable/10.1086/345785.
Plummer, Martyn. 2015. “JAGS Version 4.0.1 User Manual.” http://sourceforge.net/projects/mcmc-jags/files/Manuals/4.x/.
R Core Team. 2016. “R: A Language and Environment for Statistical Computing.” Vienna, Austria: R Foundation for Statistical Computing. https://www.R-project.org/.
Su, Zhenming, Milo D. Adkison, and Benjamin W. Van Alen. 2001. “A Hierarchical Bayesian Model for Estimating Historical Salmon Escapement and Escapement Timing.” Canadian Journal of Fisheries and Aquatic Sciences 58 (8): 1648–62. https://doi.org/10.1139/cjfas-58-8-1648.
Thorley, J. L. 2013. “Jaggernaut: An R Package to Facilitate Bayesian Analyses Using JAGS (Just Another Gibbs Sampler).” Nelson, B.C.: Poisson Consulting Ltd. https://github.com/poissonconsulting/jaggernaut.
Wickham, Hadley. 2014. “Tidy Data.” Journal of Statistical Software 59 (10): 1–22. http://www.jstatsoft.org/v59/i10.